This forum is intended for questions about kinetics, Surface Plasmon Resonance and the instruments related to these techniques.
The impact of bulk shift on kinetic analysis
- hnnigel123
- Topic Author
- New Member
-

Less
More
- Thank you received: 0
2 years 10 months ago - 2 years 10 months ago #1
by hnnigel123
The impact of bulk shift on kinetic analysis was created by hnnigel123
Hi,
I have little problem understanding the impact of a bulk shift on kinetic analysis.
According to my latest BiaCore run, I experienced bulk shifts due to buffer mismatch. Since I know the issue, I can fix this trouble easily. But I have a limited amount of analyte, so I want to find out whether the bulk shift is affecting the kinetic analysis and understand how the bulk shift affects a kinetic analysis.
So my binding curves have bulk shifts, which are about 20 % of the binding response from each concentration of the analyte. And I have used a 1:1 binding model to fit it, and it gave reasonable ka, kd and KD values. But of course, I have got a comment from analysis saying "bulk shift detected".
Seems like the bulk shift in my run affected the measured Rmax value in the analysis. From a topic in the SPR-PAGE forum, Arnoud replied that the Rmax does not affect the ka and kd. But some literature showed that the ka and Rmax are related (please correct me if I am wrong). And this makes me a bit nervous about my results.
Does bulk shift affect the kinetic analysis? If it does, then how are they affect the kinetics? And are there any evidences that Rmax is not related to ka and kd analysis?
Thank you,
hnnigel123
I have little problem understanding the impact of a bulk shift on kinetic analysis.
According to my latest BiaCore run, I experienced bulk shifts due to buffer mismatch. Since I know the issue, I can fix this trouble easily. But I have a limited amount of analyte, so I want to find out whether the bulk shift is affecting the kinetic analysis and understand how the bulk shift affects a kinetic analysis.
So my binding curves have bulk shifts, which are about 20 % of the binding response from each concentration of the analyte. And I have used a 1:1 binding model to fit it, and it gave reasonable ka, kd and KD values. But of course, I have got a comment from analysis saying "bulk shift detected".
Seems like the bulk shift in my run affected the measured Rmax value in the analysis. From a topic in the SPR-PAGE forum, Arnoud replied that the Rmax does not affect the ka and kd. But some literature showed that the ka and Rmax are related (please correct me if I am wrong). And this makes me a bit nervous about my results.
Does bulk shift affect the kinetic analysis? If it does, then how are they affect the kinetics? And are there any evidences that Rmax is not related to ka and kd analysis?
Thank you,
hnnigel123
Last edit: 2 years 10 months ago by hnnigel123.
Please Log in or Create an account to join the conversation.
- Arnoud
- Moderator
-

Less
More
- Thank you received: 0
2 years 10 months ago - 2 years 10 months ago #2
by Arnoud
Replied by Arnoud on topic The impact of bulk shift on kinetic analysis
Interesting question because most evaluation programs calculate automatically the bulk shift (RI) during fitting. First let us look at the theoretical side.
The total response at a certain time during a kinetic run is the result of the association + dissociation (the kinetics) and the bulk contribution. The bulk contribution is assumed constant during the association and absent during the dissociation. The Rmax is dependent on the number of ligand sites, analyte concentration and the ligand to analyte size ratio ( www.sprpages.nl/sensorgram-tutorial/a-curve ). When there is a bulk shift, this will make the Rmax higher but will not change the kinetics.
In practise the method of double referencing is capable to compensate for direct bulk shift differences by subtracting the reference channel from the active channel and then subtracting the zero-analyte concentration from all other curves to compensate for volume displacement differences between reference and active channel ( www.sprpages.nl/experiments/data-processing ).
This brings us to the real-world fittings. The fitting programs in general try to fit ka, kd, Rmax, RI and km (mass transport) in one go. Most of the times this goes well but in others the math gets stuck in a local minimum ( www.sprpages.nl/data-fitting/theory ) and the fitting is not as good as it can be. Close inspection of the fitting will reveal high residuals. Therefore, always do several fittings with different initial values to get some idea about the best fit.
Because the model does not know is programmed to vary the parameters and to minimize the Chi2 it can be that a high calculated RI value will fit the results or a high km. this means that all reported values should be examined carefully.
The report function of the Biacore BiaEvaluation will warn you for some problems and in your case the RI.
I like to start with a simple fitting, without mass transfer and bulk shift. You can add a model to the BiaEvaluation your self for convenience. Then use the initial values obtained and add bulk shift and later the mass transfer to refine the fit.
So far, the first part of your question.
Second part later..
Kind regards
Arnoud
The total response at a certain time during a kinetic run is the result of the association + dissociation (the kinetics) and the bulk contribution. The bulk contribution is assumed constant during the association and absent during the dissociation. The Rmax is dependent on the number of ligand sites, analyte concentration and the ligand to analyte size ratio ( www.sprpages.nl/sensorgram-tutorial/a-curve ). When there is a bulk shift, this will make the Rmax higher but will not change the kinetics.
In practise the method of double referencing is capable to compensate for direct bulk shift differences by subtracting the reference channel from the active channel and then subtracting the zero-analyte concentration from all other curves to compensate for volume displacement differences between reference and active channel ( www.sprpages.nl/experiments/data-processing ).
This brings us to the real-world fittings. The fitting programs in general try to fit ka, kd, Rmax, RI and km (mass transport) in one go. Most of the times this goes well but in others the math gets stuck in a local minimum ( www.sprpages.nl/data-fitting/theory ) and the fitting is not as good as it can be. Close inspection of the fitting will reveal high residuals. Therefore, always do several fittings with different initial values to get some idea about the best fit.
Because the model does not know is programmed to vary the parameters and to minimize the Chi2 it can be that a high calculated RI value will fit the results or a high km. this means that all reported values should be examined carefully.
The report function of the Biacore BiaEvaluation will warn you for some problems and in your case the RI.
I like to start with a simple fitting, without mass transfer and bulk shift. You can add a model to the BiaEvaluation your self for convenience. Then use the initial values obtained and add bulk shift and later the mass transfer to refine the fit.
So far, the first part of your question.
Second part later..
Kind regards
Arnoud
Last edit: 2 years 10 months ago by Arnoud.
Please Log in or Create an account to join the conversation.
- Arnoud
- Moderator
-

Less
More
- Thank you received: 0
2 years 10 months ago - 2 years 9 months ago #3
by Arnoud
Replied by Arnoud on topic The impact of bulk shift on kinetic analysis
Second part:
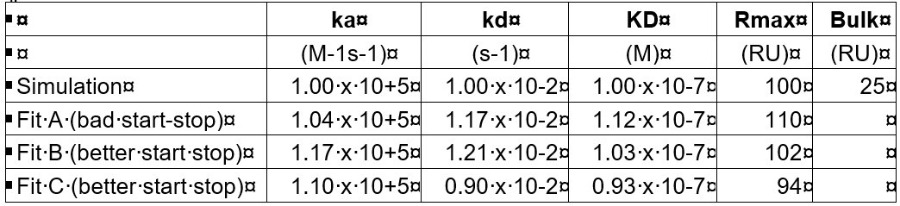
I did some simulations withmass-transfer bulk jumps and then fitted the curves. In general, the original values were resolved quite good. The placing of the start and end of the association part can influence the result a little. To give you an idea:
The fittings were done on very clean data (no noise, no drift or non-specific binding) so real-world fittings will be more difficult. I hope this will bring some more confidence in your results.
Kind regards
Arnoud
I did some simulations with
The fittings were done on very clean data (no noise, no drift or non-specific binding) so real-world fittings will be more difficult. I hope this will bring some more confidence in your results.
Kind regards
Arnoud
Last edit: 2 years 9 months ago by Arnoud. Reason: see crossed words
Please Log in or Create an account to join the conversation.
- hnnigel123
- Topic Author
- New Member
-

Less
More
- Thank you received: 0
2 years 9 months ago #4
by hnnigel123
Replied by hnnigel123 on topic The impact of bulk shift on kinetic analysis
Hi Arnoud,
Sorry about the late reply!
Thank you for doing the simulations. Your simulation makes me more confident with my results
I also have played around with the analysis software. With the constant RI value, the 1:1 model struggled to fit the curves (maybe because I had 5 different concentrations (very low stock concentration), and each concentration had different RI values). But eventually, it helped me to understand that the RI mostly affects the Rmax value but not ka, kd and KD values.
Now I have other problems (from a separate project!), probably I will chat in the forum soon!
Thank you a lot!
Regards,
hnnigel123
Sorry about the late reply!
Thank you for doing the simulations. Your simulation makes me more confident with my results
I also have played around with the analysis software. With the constant RI value, the 1:1 model struggled to fit the curves (maybe because I had 5 different concentrations (very low stock concentration), and each concentration had different RI values). But eventually, it helped me to understand that the RI mostly affects the Rmax value but not ka, kd and KD values.
Now I have other problems (from a separate project!), probably I will chat in the forum soon!
Thank you a lot!
Regards,
hnnigel123
Please Log in or Create an account to join the conversation.
Moderators: Arnoud, Arnoud